you may also like

makeadent


Radix Labs


Could your lab do more for you?


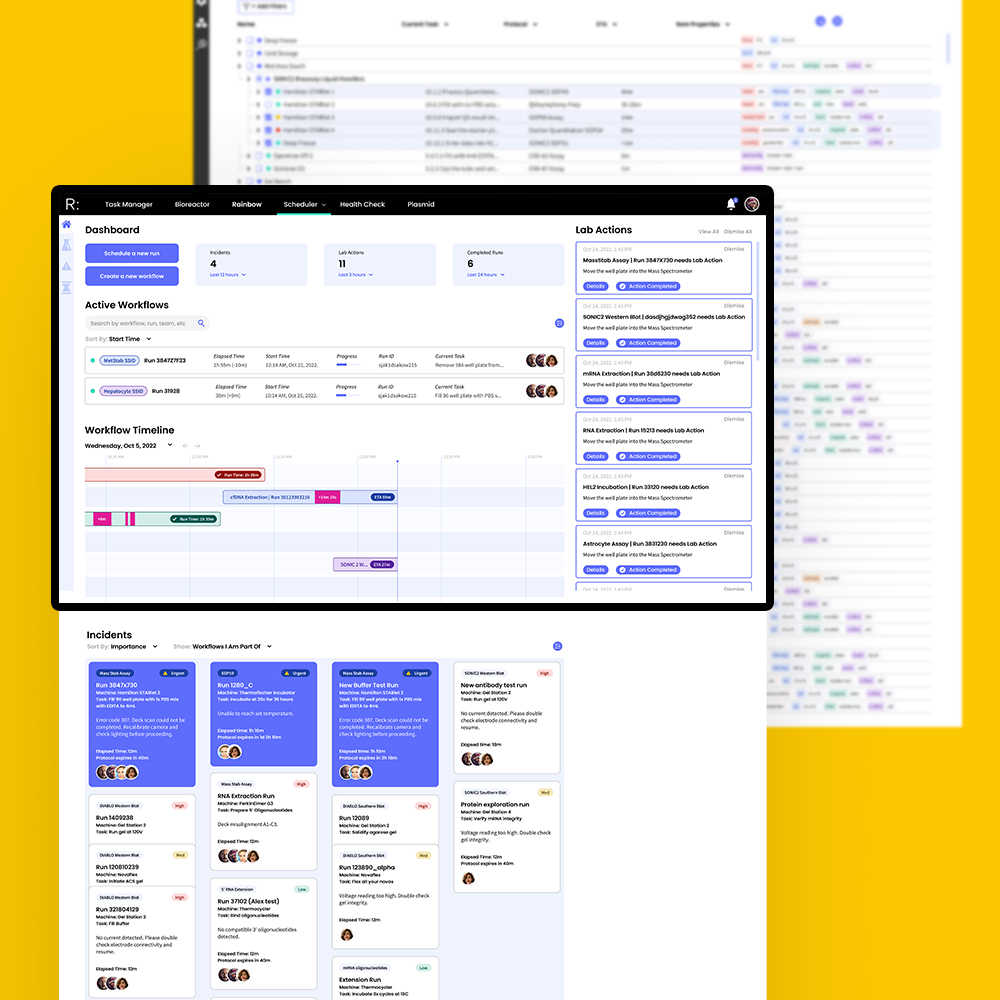
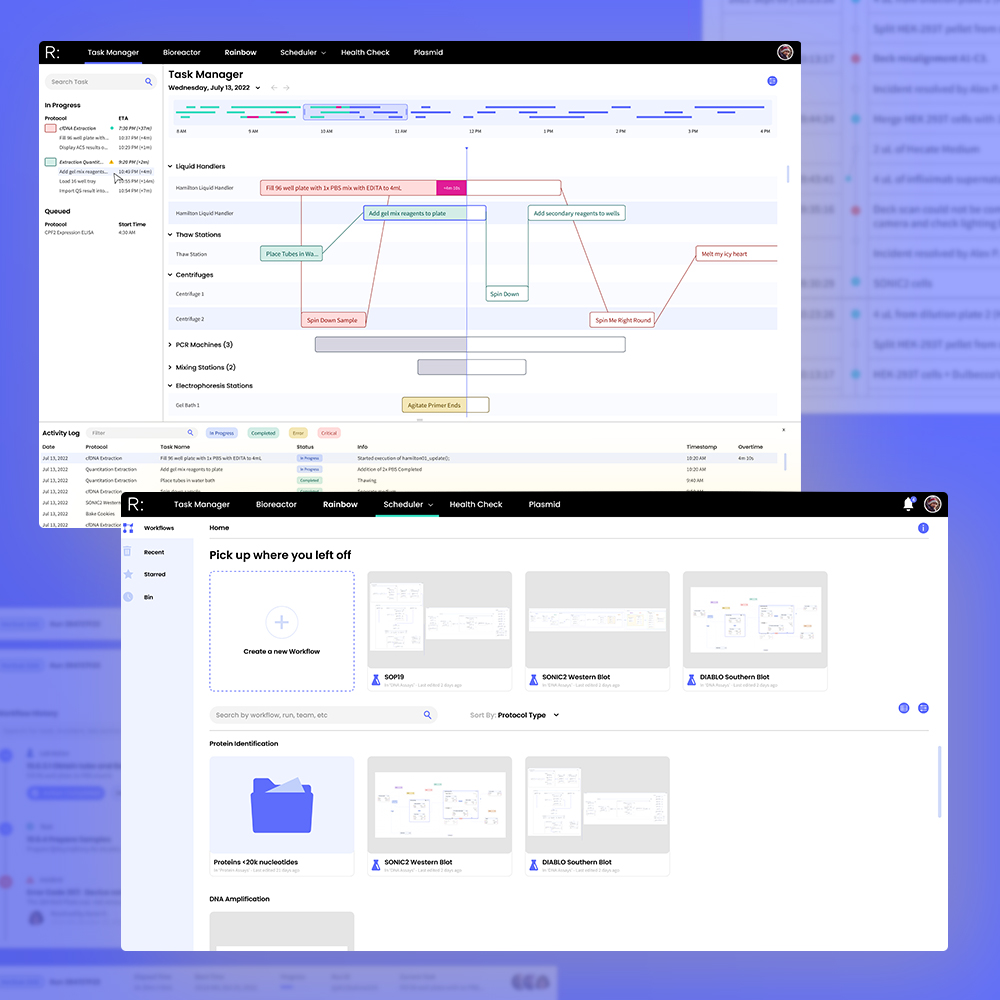
Working together with the engineering team, I was tasked with designing a platform that could translate and automate complex lab procedures into machine protocols while providing a clear, real-time overview of the current state of the lab.
I began the research phase by interviewing stakeholders to create user personas and journey maps to establish an information hierarchy. How would students, junior and senior scientists, lab technicians, and supervisors interact with the scheduler differently? Each group had distinct priorities when designing protocols, scheduling workflows, monitoring data, managing equipment, resolving conflicts, and troubleshooting. The final product needed to bring all these needs together into a single, intuitive interface.
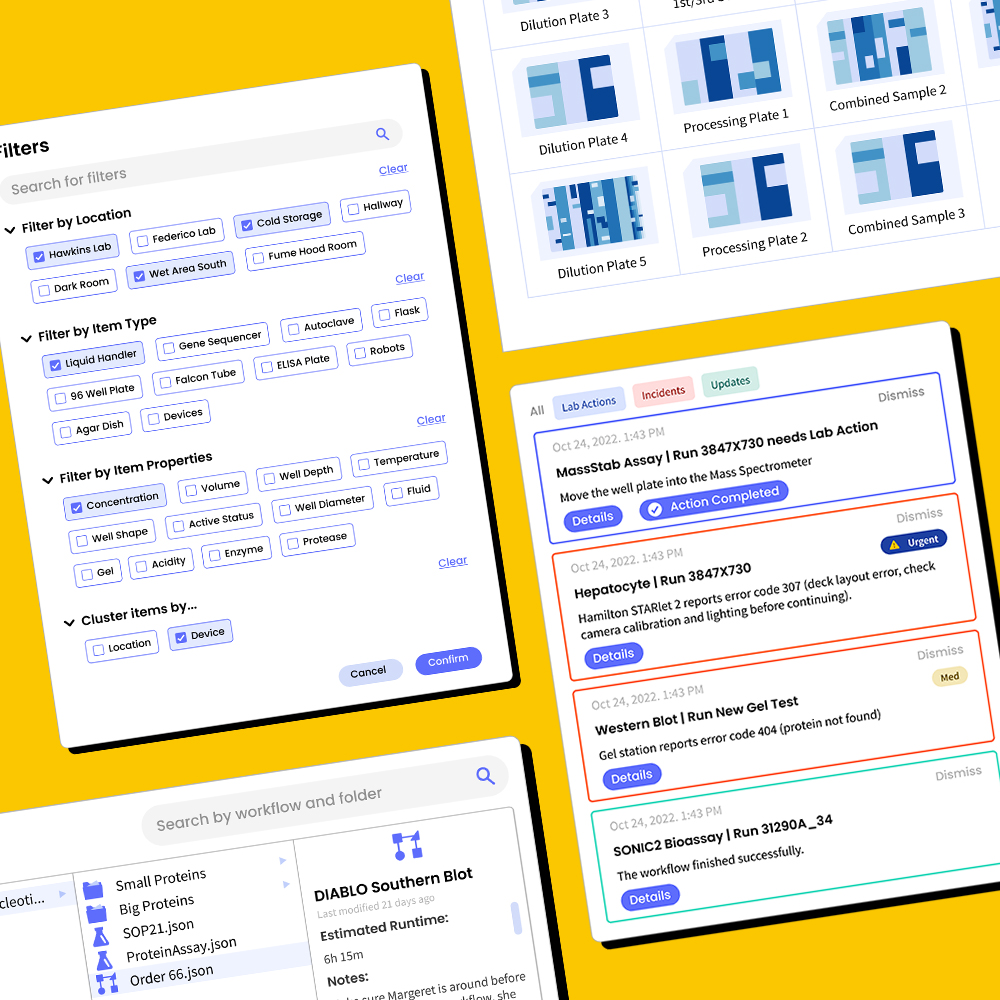
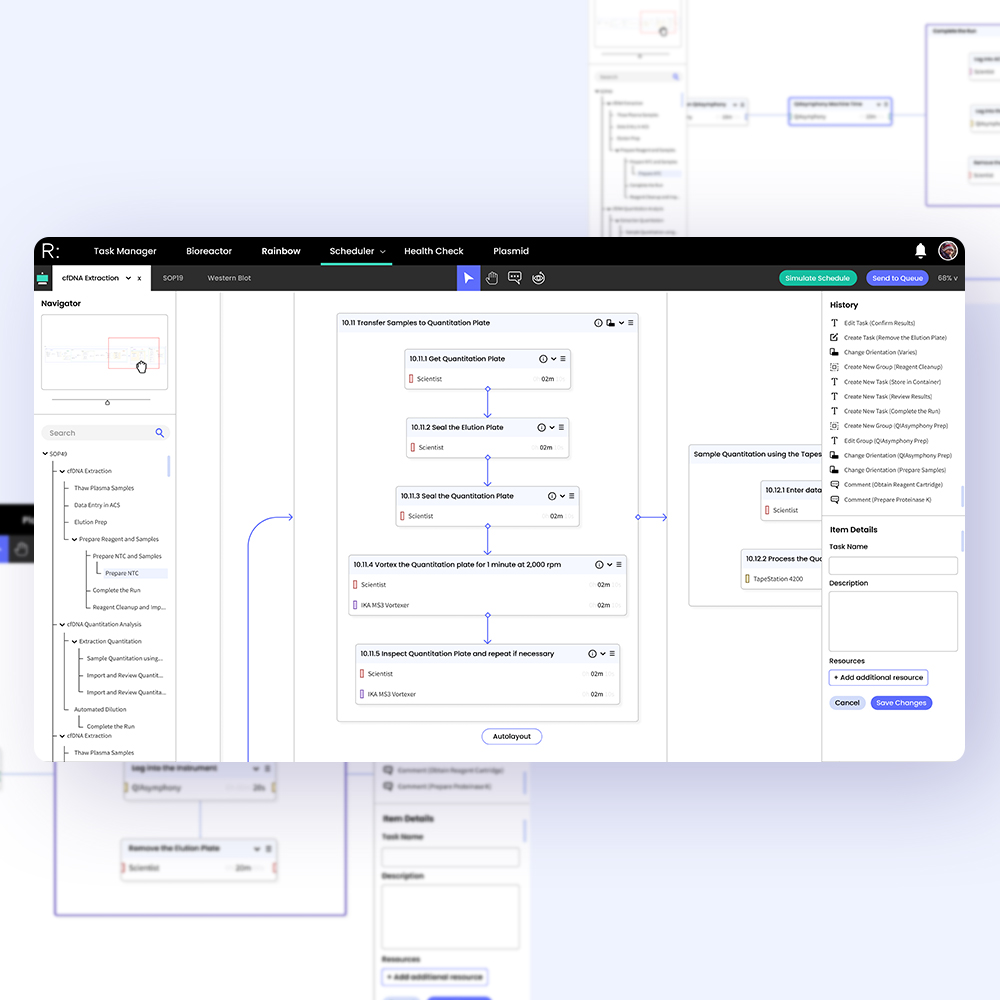
I designed and tested workflows for core tasks including troubleshooting incidents, processing lab actions, onboarding lab equipment, filtering large (>10,000 item) datasets, accessing protocol histories, expanding well plates, and simulating protocol schedules before committing.
To reduce cognitive load, I used visual representations wherever possible while maintaining the flexibility for advanced, scalar queries. Onboarding actions were simplified through an ID3-style string naming system for precise bulk processing.
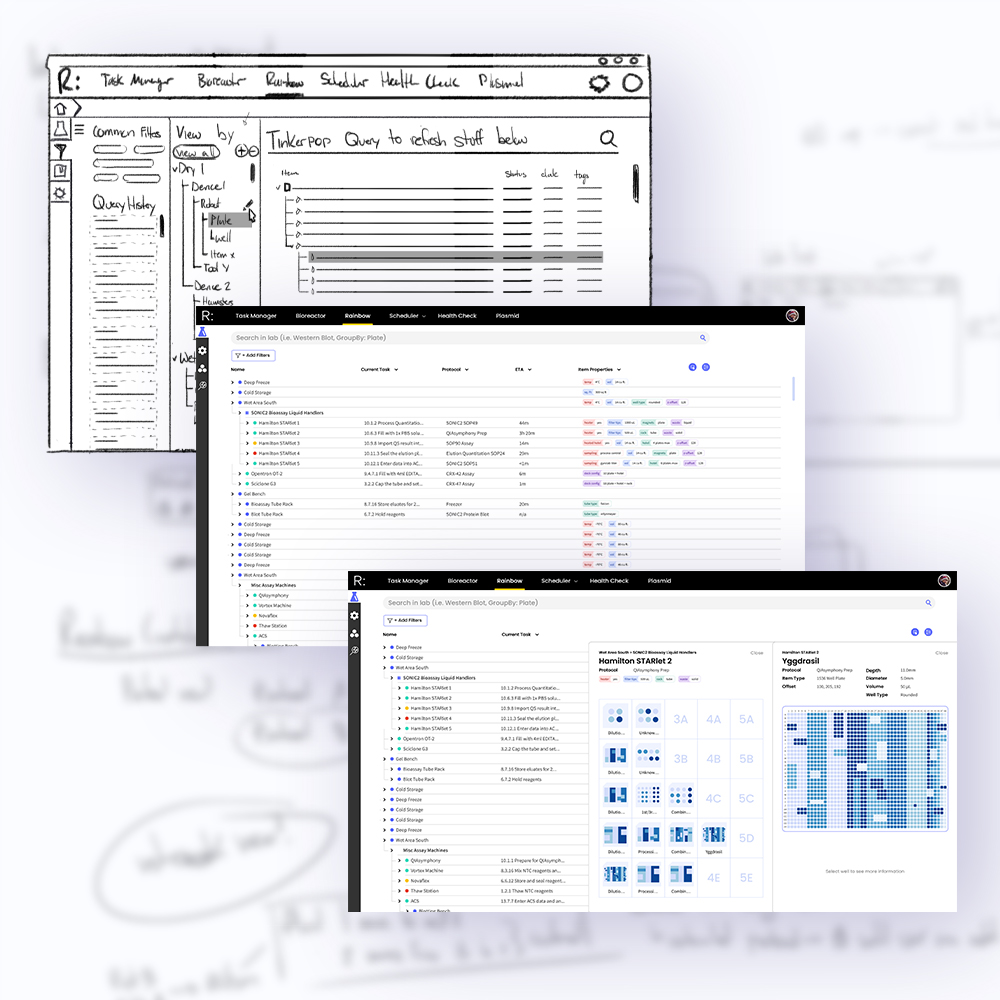
Iterative prototypes, progressing from low-fidelity sketches to high-fidelity Figma builds. This provided visual structure for the engineering team and also enabled us to conduct live feedback sessions with clients to better understand user heatmaps and improve clarity of communication and accessibility.